Discoveries In Microbiology 2019: The influence of microscopic organisms such as bacteria and viruses on our everyday life has been underestimated for a long time.
In the recent decade, research has shown how bacteria and viruses evolve, attack, and defend against each other, as well as influence essential processes on Earth.
Research performed in 2019 has uncovered a lot of new evidence in several areas of microbiology and virology that contributes to this changed view of microscopic organisms.
For instance, studies show how bacteria may influence climate change and protect mammals from certain viruses. We now know that bacteria are capable of complex reactions as a group, as seen in an original study on P. aeruginosa.
Our understanding of viruses has dramatically improved, too. This understanding is not limited to human viruses such as HIV – we now understand bacteriophages a little better, which is vital in the world where antibiotic resistance continues to grow.
Latest Discoveries in Microbiology 2019
We hope our list of the most exciting news in 2019 would showcase this progress. Here are the latest research in medical microbiology.
A blast from the past: viruses that infect an ancient group of archaea was found to have unique fusion proteins [Finland, February 2019]
Viruses can get inside the cells due to their ability to integrate the cellular membrane proteins into their protein envelopes. This integration is made possible due to a unique protein group called membrane fusion proteins.
Fusion proteins of the viruses that infect eukaryotic organisms, including mammals, are well-studied. Viruses that specialize in prokaryotes are being investigated only recently in this regard, and not much is know about their proteins.
A team of researchers from the University of Helsinki has studied several haloarchaeal pleomorphic viruses pleomorphic viruses(HRPVs) that infect a group of prokaryotic organisms – archaea, specifically those who thrive in environments with high salt content. Their research has revealed that:
- The envelopes of HRPVs are decorated with spikes that supposedly assist in attachment and invasion of cells.
- An analysis of the spikes in two related HRPV viruses has revealed a common spike protein, VP5.
- VP5 of those proteins had a mass of 57kDA and also had a V-fold that was absent in the fusion proteins of other viruses.
- VP5 was proven to be necessary for fusion of the viral envelope with the host membrane.
Up to date, only three groups of viral fusion proteins have been discovered. The new protein found in HRPV viruses belongs to a new, fourth group, as it has its own unique features not shared with the other three fusion protein types.
![]()
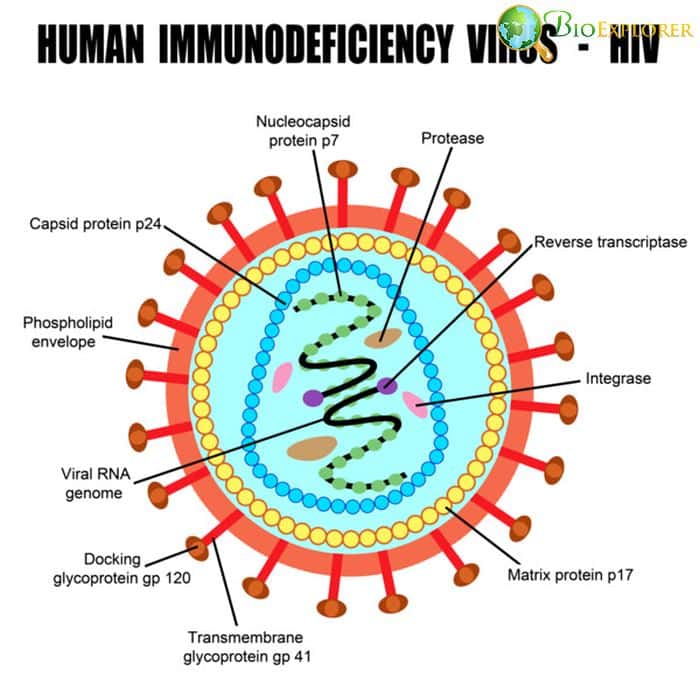
HIV can hijack immune cells: a complex mechanism that allows HIV to use macrophages to invade other immune cells of the body found [USA, December 2019]
The main danger of HIV infection lies in the fact that this virus infects immune cells – mainly T helper cells, or CD4 cells. The infection is further complicated by the fact that the virus can “hide” in other immune cells, such as dendritic cells that reside in the lymph nodes instead of the bloodstream.
It was still unknown how the HIV virus actually gets inside those cells initially. A research team from the National Institute of Allergy and Infectious Diseases (NIAID) decided to look into the early events during HIV infection in mice. They have found out that:
- As HIV enters the bloodstream, it uses a type of lymphoid macrophages as “transport ” that takes the particles to lymph nodes.
- These macrophages are called sinus lining macrophages and possess a protein called MFG-E8.
- MFG-E8 can link to the viral particles so that the virus can enter the macrophages.
- Afterward, the viral particles are passed to the dendritic cells in the lymph nodes through a protein called α vß-integrin.
- If the MSG-E8 protein or the integrin protein is absent or blocked, the virus cannot be spread to dendritic cells.
These new findings are precious for future HIV research. However, it is not clear yet whether this mechanism is more advantageous for the virus itself or is actually a mechanism of protecting the host.
![]()
How Shigella bacterium becomes an immortal Highlander: the researchers have found a mechanism that helps a bacterium with blocking cell death [Germany, December 2019]

Shigella is a bacterial species that gets inside the cells of the intestinal epithelium and can cause debilitating diarrhea. It has multiple mechanisms aimed at evading the immune system. A team of German researchers has found another one-that makes the bacterium withstand the order to die:
- The immune system can produce several enzymes that trigger cell death in the infected cells.
- It was shown in a mouse model of Shigella infection, that these bacteria can use specific agents to block these compounds.
- These compounds are called lipopolysaccharides (LPS) and are located on the bacterial cell surface.
- Bacterial LPS binds the enzymes that trigger cell death in the cell called caspases.
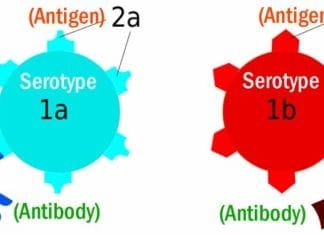
- This is a special kind of LPS that contains a structure called O antigen.
- If the Shigella strain lacks an LPS with an O antigen, such bacteria cannot block caspases and are vulnerable to the effect of the immune system.
Discover The Differences Between Antigen And Antibody
This information can be crucial in understanding Shigella infection. It can be potentially used in developing drugs that would counteract the problem.
![]()
Catching the bacterial spy: the researchers have found a protein that helps resistant bacteria sense antibiotics [USA, September 2019]
Many infections that can be potentially caught at hospitals can be tough to treat because these bacteria are highly resistant to antibiotics. One such pathogen is vancomycin-resistant enterococci (VRE), for instance, Enterococcus faecalis.
In normal conditions, enterococci are a part of healthy flora. In the right circumstances, they can also potentially evolve into dangerous pathogens resistant to common antibiotics. A team from the University of Texas Health has been researching how this species evades the effect of antimicrobial peptides:
- It was known that the Enterococcus faecalis can become resistant to antibiotic daptomycine and antimicrobial peptides produced by the immune system.
- The bacteria managed to evade death caused by these agents by changing the structure of its cellular membrane.
- The researchers have discovered that this bacterium produces a particular compound, called LiaX.
- LiaX can sense the presence of antibiotics and trigger the system that causes the remodeling of cellular membrane.
If it could be possible to block LiaX, the bacteria would not be able to defend themselves against antibiotics and would not develop multiple resistance. If such a new agent were designed, it would be a breakthrough in treating hospital-acquired resistant infections.
![]()
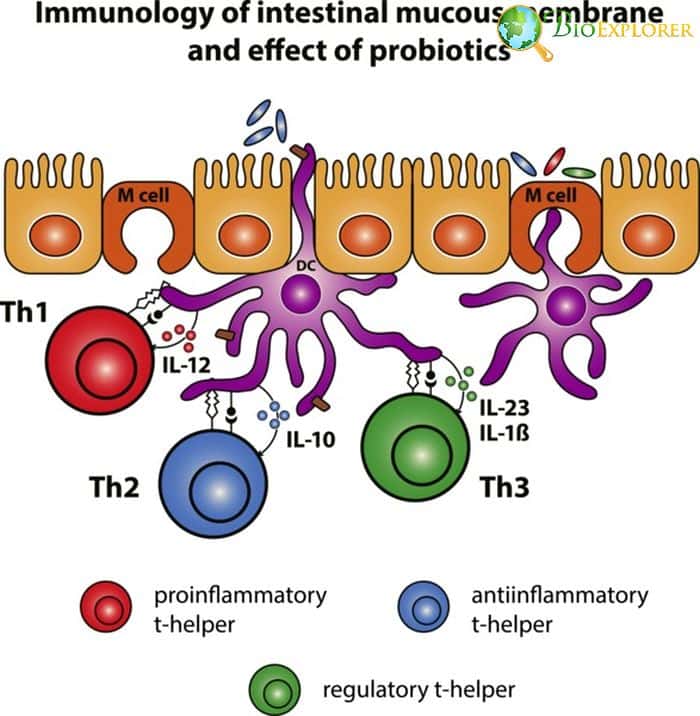
Sometimes probiotics are not safe: bacteria from probiotics capsules found in bloodstream of ICU patients [Israel-USA, November 2019]
.
Probiotics are routinely prescribed to patients that take antibiotics in order to compensate for the death of symbiotic bacteria in our guts. Still, in some cases probiotics cause more harm than good.
- Several ICU patients in Boston’s children’s hospital have developed Lactobacillus bacteremia.
- Lactobacillus is a common component of probiotic routinely prescribed to hospital patients.
- The patients received a probiotic containing Lactobacillus rhamnosus strain GG (LGG) through the feeding tube as part of the ICU treatment.
- This exact strain was later found in the blood of 6 patients in large quantities, which means the development of bacteremia.
- Genomic analysis has shown that the lactobacteria circulating in the patient’s blood were genetically identical to the strains present in the probiotic capsules.
- The only exception was the presence of an SNP in some bacteria that conferred antibiotic resistance.
This study shows that Lactobacilli can cause bacteremia in severely compromised patients and can also evolve in the bloodstream. It points to the fact it is necessary to rethink the use of probiotics in certain groups of patients.
![]()
E. coli dangerous strategy: the researchers have found the mechanism that increases E.coli virulence [USA, July 2019].
Escherichia coli is a common gut bacterium that usually does not affect our health. Besides this harmless e.coli species, there is a strain called Escherichia coli O157: H7 (EHEC) that can reside in poorly prepared and processed food and can potentially cause severe disease, with bloody diarrhea and blood in the urine.
The main danger of this bacterium is its Shiga toxin that is responsible for the disease symptoms. It is essential to understand how the virulence of this dangerous strain is regulated.
- Pathogenic and non – pathogenic E.coli both have an sRNA factor DicF, which regulates cell division and metabolism.
- The research team has compared the activity of DicF of both strains in conditions with low and high oxygen levels.
- It was found out that DicF activates in response to low oxygen conditions.
- In pathogenic E.coli, there are three additional copies of DicF, and these copies are located within areas called pathogenicity islands.
- As a result, in the pathogenic E.coli, activation of DicF in response to low oxygen levels leads to increased activity of the pathogenicity islands levels, and also to an increase in Shiga toxin production.
As it is known that pathogenic E.coli strain can travel to specific areas of the colon that have very low oxygen levels, it seems that the bacterium begins to actively divide and produce toxins as soon as it reaches its destination due to this regulatory factor. These new food microbiology news not only explain the mechanisms of the virulence of this particular strain, but they also point to a potential therapeutic target that could help with the possible treatments
![]()
Round and round the antibiotic we go: new antibiotic evasion strategy discovered in bacteria [Denmark-USA, November 2019]
Pseudomonas aeruginosa is an opportunistic species that can cause severe lung disease in patients with compromised immunity, for instance, in the case of cystic fibrosis. This species is also known for its quorum sensing behavior – the ability to react to specific signals together as a community.
A team of scientists has tried to create a model that would reflect the behavior of these bacteria on the lung surface in Petri dishes
- The bacteria were placed in Petri dishes and cultures in a way that resembled colonization of mucosal surfaces of the lungs.
- The bacteria were challenged with the presence of antibiotics or bacteriophages.
- It was established that in response to the bacteriophage infection or addition of antibiotics, the bacteria produce a particular factor called PQS.
- In response to PQS, the whole bacterial community reorganizes to evade the harmful agent and “swims around” dangerous areas in the colonies.
As the danger of P. aeruginosa infection grows, and this species is highly antibiotic-resistant, it is essential to understand the factors that the organism uses to protect itself to potentially block them.
![]()
The loop of heat and doom: as the planet heats up, bacteria may accelerate climate change, even more, study finds [UK, November 2019]

A team of researchers has decided to study how bacteria adapt to the changes in global temperatures. It was found that:
- Bacteria that are adapted to higher temperatures do not significantly change the speed of their growth, with the temperatures increase.
- Bacteria that are used to milder temperatures lower than 45C adapt to the temperature shift and accelerate their growth.
- The bacteria in this range also produce more carbon dioxide as a result.
- A mathematical model developed by the research team shows that the rise in temperatures would affect these middle-range bacteria and cause an increase in carbon dioxide levels produced by them.
According to the study, global warming would trigger a positive loop in which the temperature increase would also accelerate carbon dioxide synthesis, which, in turn, would also contribute to warming. This is a new, concerning factor that should be taken into account in modeling the consequences of climate change
![]()
Gut microbes as hidden knights: gut microbiota was shown to protect their host from rotavirus infection [USA, October 2019]
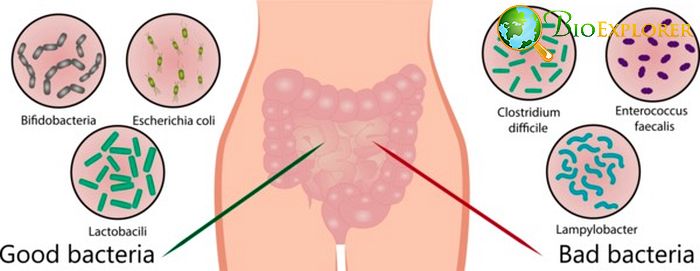
Rotavirus infection is routinely encountered by both adults and children. It usually causes significant vomiting and diarrhea and can be deadly in younger children and infants. Currently, it is recommended to vaccinate infants against rotavirus to prevent such a risk.
A research team in Institute for Biomedical Sciences at Georgia State University was studying rotavirus infection in mice and has come across unique event:
- The team has accidentally created a line of immunodeficient mice and rotavirus resistant at the same time.
- The study of the gut microbiota in the rotavirus resistant mice has shown that these mice had a high quantity of specific bacteria types called segmented filamentous bacteria (SFB).
- The presence of SFB made mice resistant to rotavirus.
- Rotavirus sensitivity in other mice types can be corrected by a fecal transplant containing SFB.
- SFB was shown to influence the production of several immune proteins and cause a high turnover of epithelial cells in the gut, preventing the rotavirus from infecting them.
- If the rotavirus is incubated together with feces containing SFB, it can be directly neutralized.
This discovery at present cannot directly help in rotavirus infection treatment in humans. Still, it can help explain why certain people are susceptible to this virus, while others are not. And the answer may lie in the microbiota composition.
![]()
Sending saboteurs from hiding: secrets of Leishmania viruses uncovered [Canada-Japan, July 2019]
Leishmania is a microscopic parasite that can reside within our cells. It can into the bloodstream with a bite of sand fly. After that, Leishmania cells get inside the host cell and hides in a special vacuole to evade cell’s defenses.
Despite being in hiding, this parasite can still release special factors that subvert the inner workings of the cell in a manner useful to the newcomer. An international team led by Professor Albert Descoteaux has investigated how exactly this organism managed to send its factors to change the processes in the host cell.
- It was found that Leishmania releases its virulence factors – lipophosphoglycan and the GP63 metalloprotease – immediately after they are engulfing by immune cells – macrophages.
- For this, Leishmania uses the cell’s own mechanisms usually employed for transport of various substances.
- Two compounds, Sec22b and syntaxin-5 that participate in the fusion of the membranes within the cell play a crucial role in this process.
- The factors end up in the crucial region of the cell – endoplasmic reticulum or ER.
- ER is connected to both the nucleus of the cell and other cell structures, which promotes spreading of the harmful substances that sabotage the cell.
- If the cell is exposed to agents blocking communication between ER and other areas, the harmful effects of Leishmania “spies ” can be mitigated.
Most of the work devoted to leischmaniasis focuses on the effects of the virulence factors themselves. This is a first study that actually describes how the virulence factors get to the target structures within the cell.
Hopefully, it would provide some ideas for novel treatments of this parasitic disease in the future.
![]()
We could not cover all the exciting news in Microbiology uncovered in 2019.For instance, researchers have discovered an incredible diversity of viruses in the least likely of places – Arctic ocean.
New roles of bacteria in the sea were uncovered – for instance, their ability to digest sugars made by algae. Novel methods allow better tracking of microbial communities and more in-depth understanding of gut microbiota.
Another research team has made a completely new and unique model that depicts how bacteria move on surfaces. Another interesting study has discovered how bacteria Stenotrophomonas maltophilia kills off its competitors.
A complex world of interactions in the microcosm of bacteria, virus, and other single-cell organisms is slowly becoming less of a mystery.
Hopefully, we would also be able to fight the dangers that emerge from it better as well.
![]()